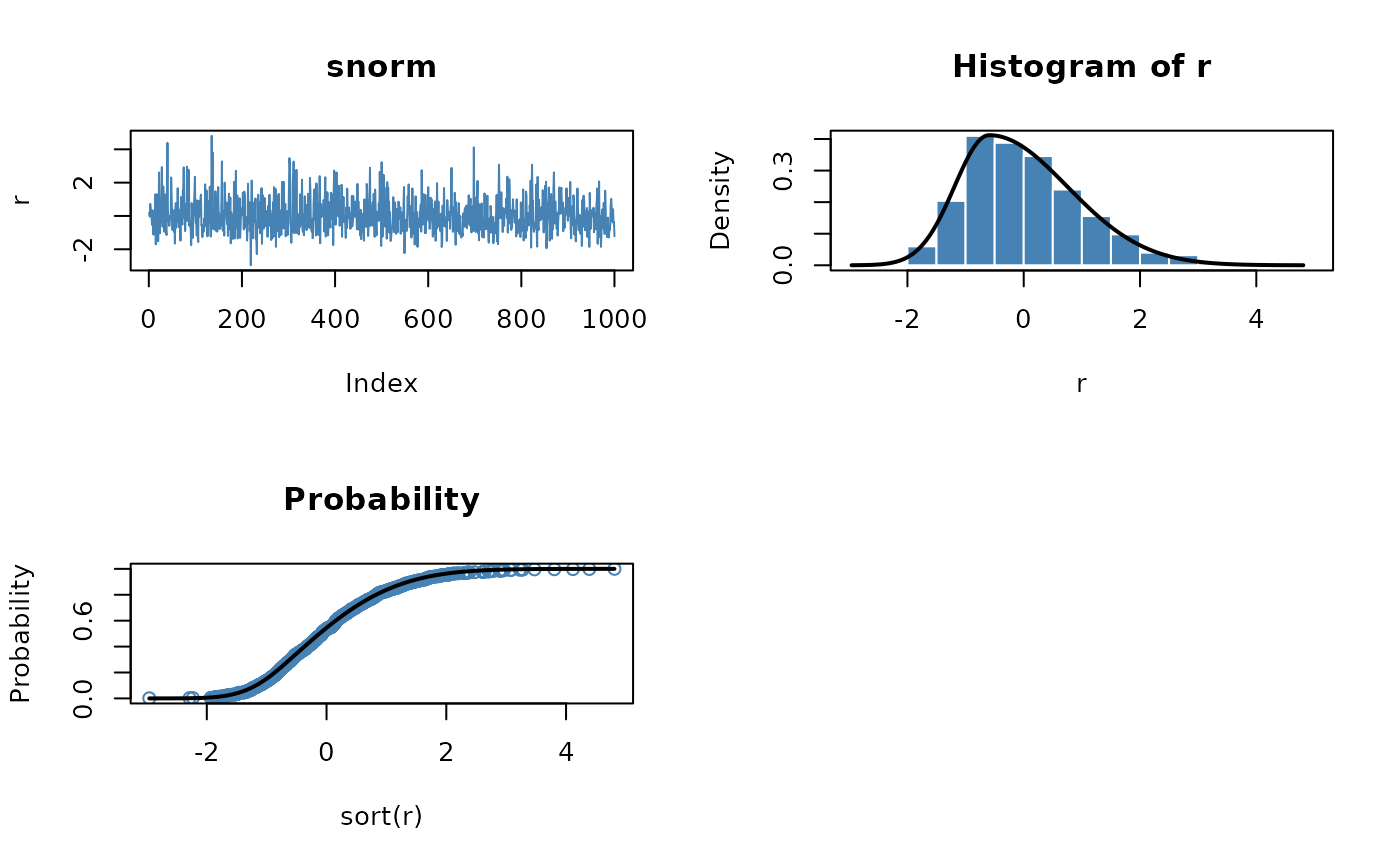Skew normal distribution
dist-snorm.RdFunctions to compute density, distribution function, quantile function and to generate random variates for the skew normal distribution. Note that there are different ways to define a skew normal distribution, check section Details to see if this is the distribution you want.
Usage
dsnorm(x, mean = 0, sd = 1, xi = 1.5, log = FALSE)
psnorm(q, mean = 0, sd = 1, xi = 1.5)
qsnorm(p, mean = 0, sd = 1, xi = 1.5)
rsnorm(n, mean = 0, sd = 1, xi = 1.5)Arguments
- x, q
a numeric vector of quantiles.
- p
a numeric vector of probabilities.
- n
the number of observations.
- mean
location parameter.
- sd
scale parameter.
- xi
skewness parameter, a positive number.
xi = 1gives a symmetric distribution (here normal).- log
a logical; if TRUE, densities are given as log densities.
Details
Skewness is based on the scheme by Fernandez and Steel (2000) but with
different parameterisation as discussed by Wuertz et al (????), in
that the skewness (xi),
mean (mean) and standard deviation (sd) are independent
parameters.
Please note that there are different ways to define a 'skew normal distribution'. In particular, the distribution discussed here is different from what is usually referred to as 'skew normal distribution' (see, for example, Azzalini 1985).
dsnorm computes the density,
psnorm the distribution function,
qsnorm the quantile function,
and
rsnorm generates random deviates.
References
Fernandez C., Steel M.F.J. (2000); On Bayesian Modelling of Fat Tails and Skewness, Preprint, 31 pages.
Wuertz D., Chalabi Y. and Luksan L. (????); Parameter estimation of ARMA models with GARCH/APARCH errors: An R and SPlus software implementation, Preprint, 41 pages, https://github.com/GeoBosh/fGarchDoc/blob/master/WurtzEtAlGarch.pdf
A. Azzalini (1985). A class of distributions which includes the normal ones. Scand. J. Statist. 12, 171-178
See also
snormFit (fit),
snormSlider (visualize),
Examples
## snorm -
# Ranbdom Numbers:
par(mfrow = c(2, 2))
set.seed(1953)
r = rsnorm(n = 1000)
plot(r, type = "l", main = "snorm", col = "steelblue")
# Plot empirical density and compare with true density:
hist(r, n = 25, probability = TRUE, border = "white", col = "steelblue")
box()
x = seq(min(r), max(r), length = 201)
lines(x, dsnorm(x), lwd = 2)
# Plot df and compare with true df:
plot(sort(r), (1:1000/1000), main = "Probability", col = "steelblue",
ylab = "Probability")
lines(x, psnorm(x), lwd = 2)
# Compute quantiles:
round(qsnorm(psnorm(q = seq(-1, 5, by = 1))), digits = 6)
#> [1] -1 0 1 2 3 4 5